Sugar modifications are not currently displayed in 3D-SNFG, although the base shape is shown. The SNFG icons are an effective method for communicating the modifications on residues. For example, the following is a heparan sulfate fragment built with the GLYCAM Webtool:
Using the 3D-SNFG icons conveys both the residue name and sulfation position.
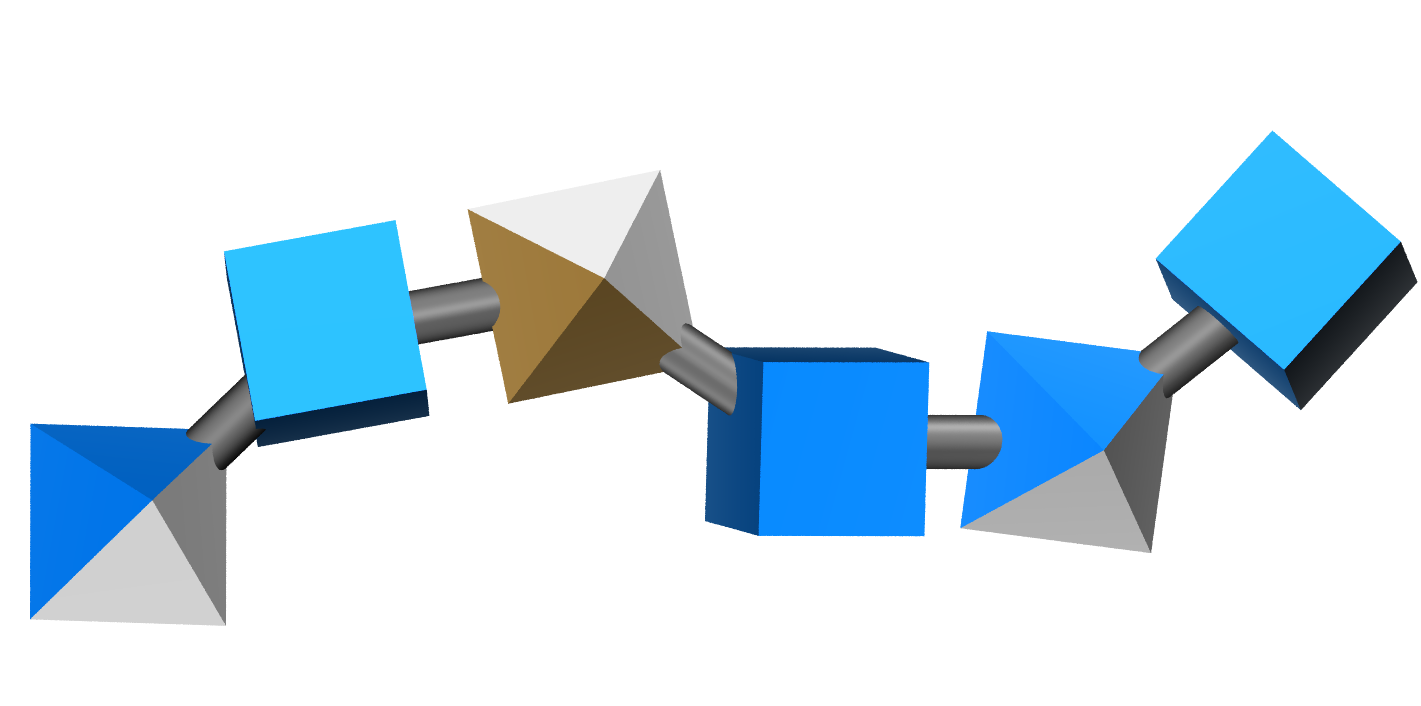
Representing the same fragment with 3D-SNFG loses the sulfation sites:
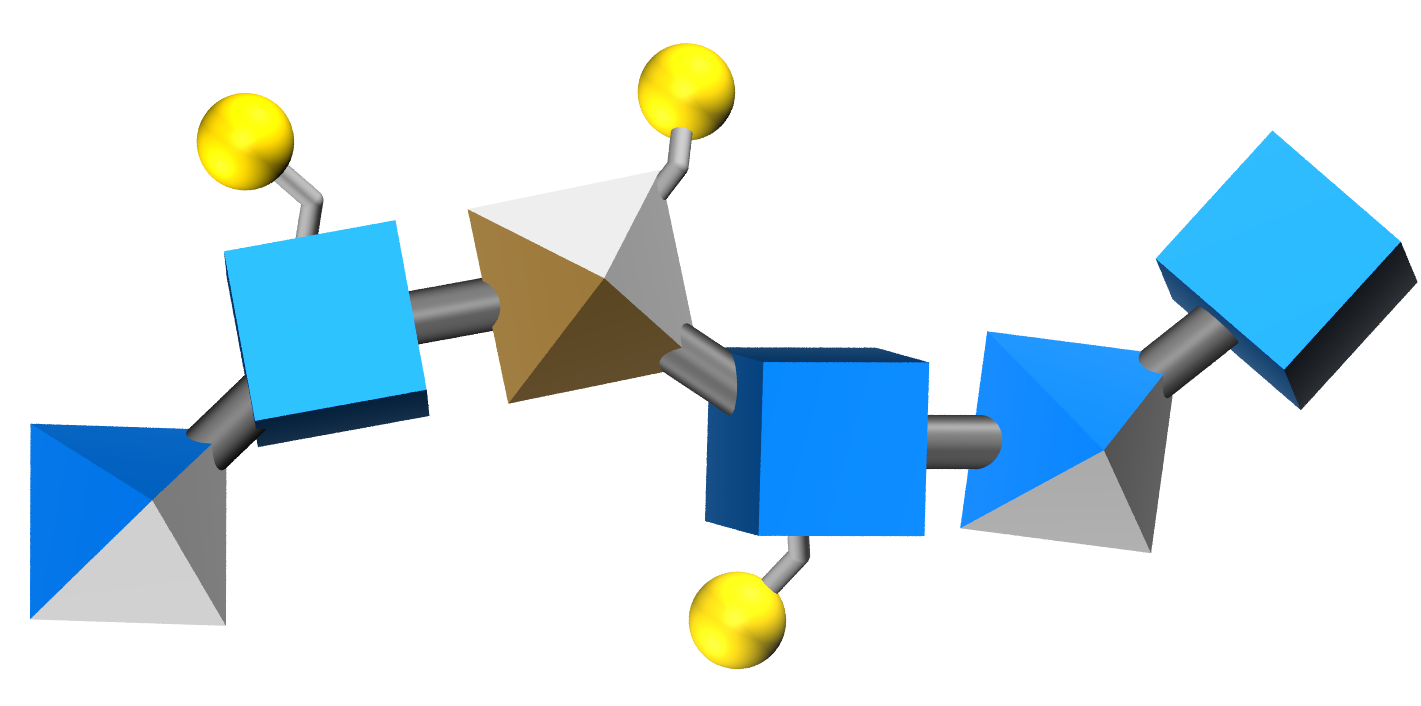
One possibility for depicting modifications with 3D-SNFG is to draw the sulfur atoms as large spheres, and the connecting atoms as silver licorice:
The sulfation position (i.e. NS, 2S) can be labeled by adding a text-box on top of the sulfur atom via any photo editing software (or PowerPoint). The drawback of this representation is that the orientation in which the molecule is viewed may obscure some sulfation sites.