This FAQ entry refers to Step 3 of the Glyoprotein Builder, where glycosylation sites are chosen for a given glycan. Here, we have uploaded 1UBQ.pdb to use as an example.
When you reach the “Attach Glycans” portion of the build, you will be presented with a list of potential glycosylation sites. They are divided into two groups, N-linking sites and O-linking sites, as shown in this image:
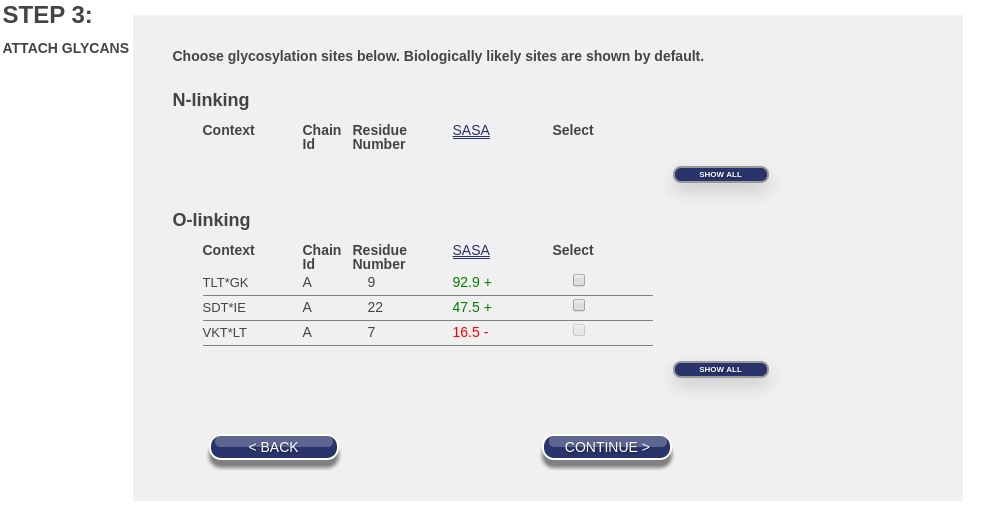
Note the buttons on the right that say “Show All”. If you click them both, the number of entries will expand as shown in this image:
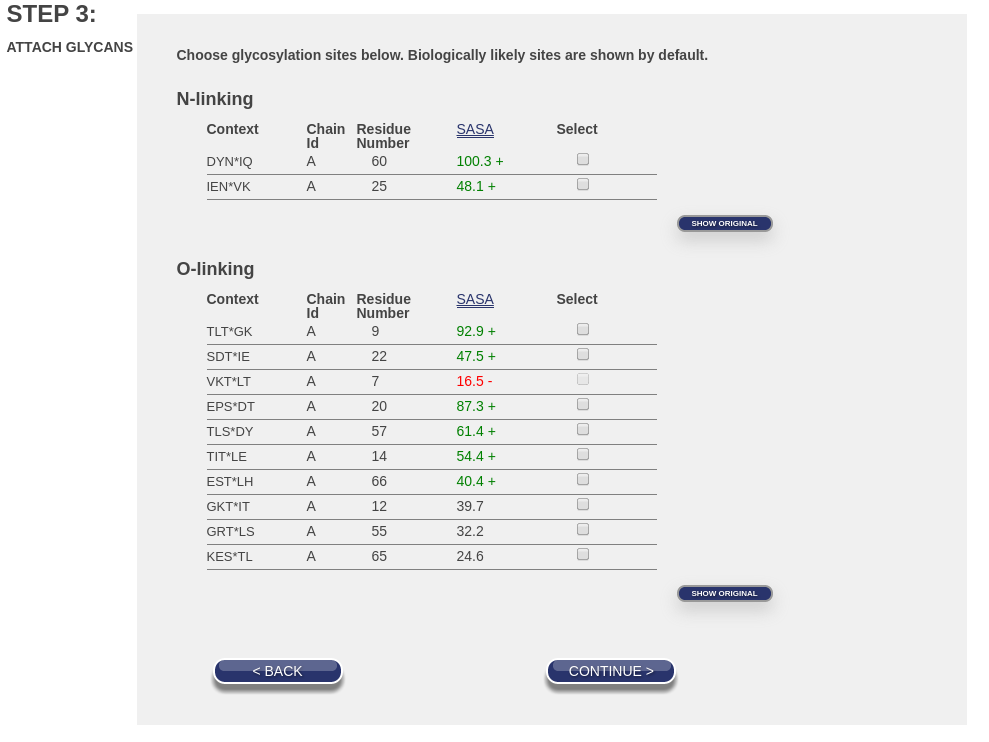
Note that the buttons now say “Show Original”. If you click them again, you will once again see a page like the first image.
In the first image, only sites that are likely to be biologically relevant are shown. However, other sites can be glycosylated, even in biological contexts, so we allow the user to choose any accessible site. To learn more about site accessibility, see this FAQ. If the user clicks “Show All”, all potential sites are shown.
We use the following methods to determine whether or not a site is likely to be biologically relevant:
N-Linking: Only sites that occur in a sequon are shown by default. A sequon is a pattern of amino acids where most N-glycosylation occurs. The pattern is ASN-X-SER/THR, where X can be anything except proline (PRO). Here, ASN stands for asparagine, SER for serine, and THR for threonine. Note the context, given in the left-hand column, in the images above. For more information, see any modern text that treats glycobiology. NCBI makes the text Essentials of Glycobiology, 2nd Ed., from 2009, publicly available. Click here to view the chapter about N-glycans.
O-Linking: Only sites identified by the NetOGlyc software (version 3.1 as of this writing) as being relevant to mucin-type O-glycosylation by GalNAc (N-acetyl-galactosamine) are shown by default. Please see the NetOGlyc site, linked earlier, for further information.
